Super-resolution microscopy reveals 3D organization of genes
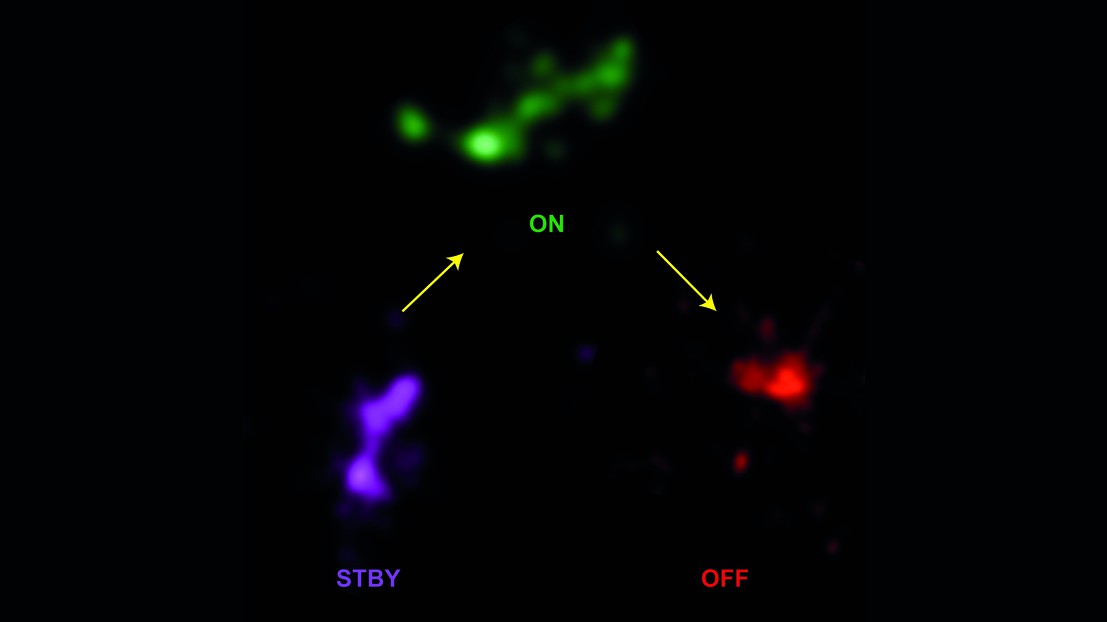
Super-resolution imaging of a DNA fragment containing nine Hox genes © Denis Duboule/EPFL
EPFL scientists have used super-resolution microscopy to study in unprecedented detail the organizational structure of genes that regulate the development of the embryo.
Image: Super-resolution imaging (STORM) of a fragment of DNA containing nine genes (Hoxd genes). The three representative shapes show the key conformations of the Hoxd gene cluster in ES cells (magenta; stand-by state), digits cells (green; active state) and brain cells (red; inactive and repressed state).
In the cell, DNA is tightly packaged in a structure called chromatin, unwinding small parts at a time to allow genes to be read and expressed. During embryonic development, a family of “architect” genes orchestrates the appearance of limbs and vertebrae, each providing precise instructions at specific times. The activity of architect genes themselves is in turn controlled by unwound DNA packaging. Taking detailed, super-resolution images of DNA, scientists from EPFL have been able to observe for the first time the nanoscale organization of architect genes. The breakthrough work is published in PNAS.
The packaging of DNA in three-dimensional space is far from random. In fact, our entire genome is highly organized to allow for tight regulation of gene expression at specific times. This spatiotemporal regulation applies to all genes – even the architect genes that control the development of the fetus. These are a family of genes called Hox, and are active during this early phase to organize the orientation of the embryo’s body, as well as limb placement and spinal development. Thousands of interactions between the Hox genes and their regulatory elements (“enhancers”) take place to achieve spatiotemporal control of their expression during embryonic development.
The labs of Denis Duboule and Suliana Manley, from EPFL’s Faculty of Life Sciences and Faculty of Basic Sciences, have now collaborated in a breakthrough study to literarily “see” how this 3D control takes place in compacted DNA. The study, led by Pierre Fabre from Duboule’s lab, looked at mouse embryonic stem cells using a technique called super-resolution microscopy, which can overcome the diffraction limit of light. Because of this, researchers can use it to study subcellular structures in greater detail than that offered by confocal microscopes.
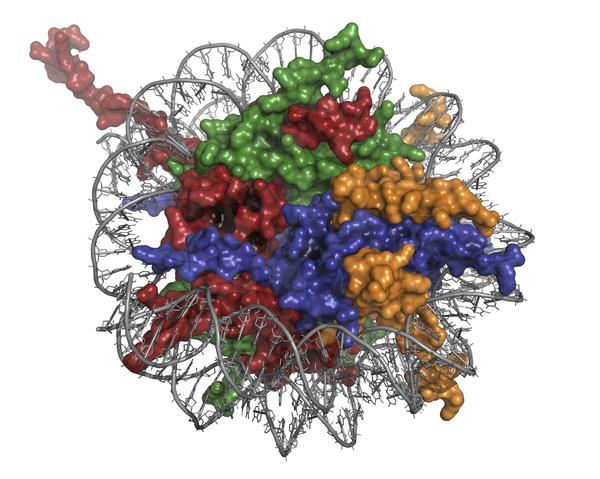 The researchers used a variant of superresolution microscopy called “stochastic optical reconstruction microscopy” or STORM, which takes high-resolution images by using sequential activation and time-resolved localization of fluorophores that can be turned on and off with light. The STORM part of the study was performed by Manley’s lab, who have longstanding expertise with it.
The researchers used a variant of superresolution microscopy called “stochastic optical reconstruction microscopy” or STORM, which takes high-resolution images by using sequential activation and time-resolved localization of fluorophores that can be turned on and off with light. The STORM part of the study was performed by Manley’s lab, who have longstanding expertise with it.
The detailed imaging in single cells revealed the compartmentalization of DNA in unprecedented detail. The researchers were able to visualize specific parts of packaged DNA called “Topologically Associating Domains” or TADs. These range from thousands to millions of DNA bases, and function as the "neighborhoods" in which the mammalian genome is divided. It is within TADs that most enhancer interactions occur to activate gene expression.
The researchers analyzed the shapes of compacted DNA in different types of mouse cells. In embryonic stem cells, they were able to observe genes decompacted before they became activated. In digit cells, the genes were turned on, and in brain cells they were completely turned off, corresponding to significantly different shapes. This confirmed that DNA organization differs between tissues that have different gene-activation profiles.
The researchers analyzed the shapes of compacted DNA in different types of mouse cells. In embryonic stem cells, they were able to observe genes before they became activated. In digit cells, the genes were turned on, and in brain cells they were completely turned off. What this confirmed was that DNA organization differs between tissues with different activation status.
The findings shed light on how gene spatial compartmentalization takes place. The arrangement allows for efficient reorganization of chromatin when transcriptional switches are activated.
Current methods can give us very limited information about the 3D organization of the genome, especially at the level of genes (kilobases) and regulatory domains (hundreds of kilobases). This has created a “gap of resolution” between the scale of the well-studied DNA double helix and the large arrangement of DNA in chromosomes.

From Marti-Renom & Mirny (2011) Bridging the Resolution Gap in Structural Modeling of 3D Genome Organization. PLoS Comput Biol 7(7): e1002125. doi:10.1371/journal.pcbi.1002125
This new understanding of DNA decompaction can now be used to study the mechanisms that open and close chromatin to regulate gene activity. “We will carry on this work using genetic strategies previously developed in the lab, while others can expand onto the rest of the genome,” says Pierre Fabre. “Seeing is believing.”
This work was funded by EPFL, the University of Geneva, the Swiss National Research Fund, the European Research Council, SystemsHox.ch, and the Claraz Foundation.
Reference
Fabre PJ, Benke A, Joye E, Nguyen Huynh TH, Manley S, Duboule D. Nanoscale spatial organization of the HoxD gene cluster in distinct transcriptional states.PNAS 26 October 2015. DOI: 10.1073/pnas.1517972112