EPFL takes part in largest study of ancient leprosy
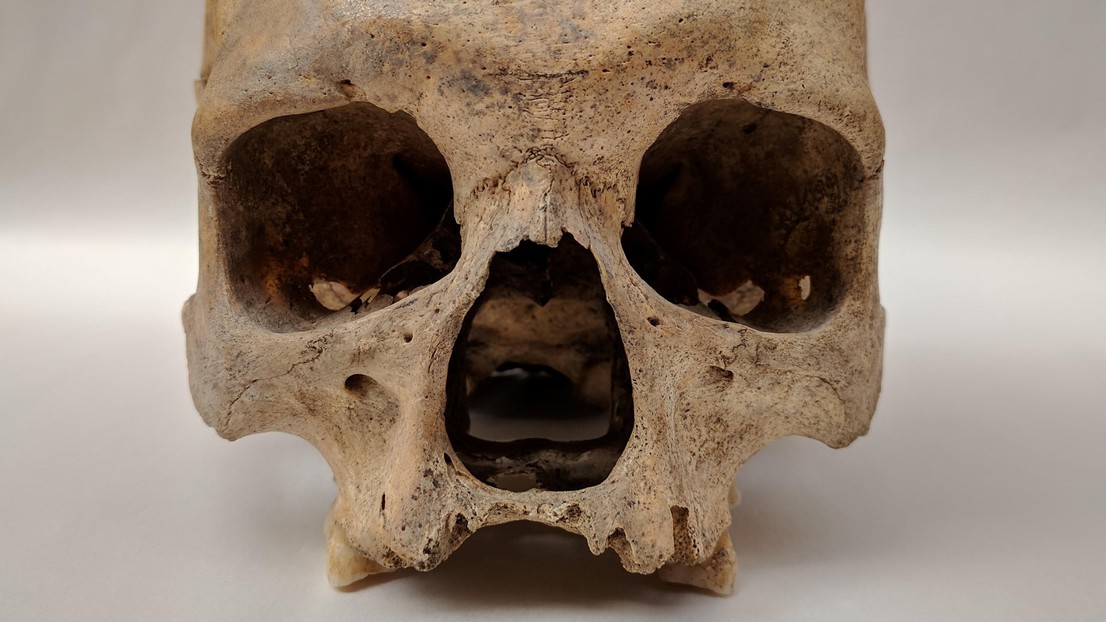
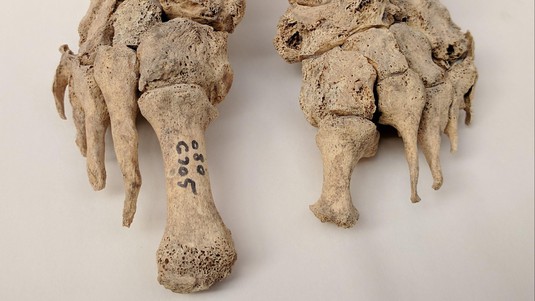
Skeletal remains showing evidence of leprosy from the Odense St. Jørgen cemetery in Denmark, which was established in 1270 and existed until 1560. Credit: Dorthe Dangvard Pedersen
The largest study to-date on the DNA of ancient leprosy bacteria reveals previously unknown diversity of strains in medieval Europe. Scientists from EPFL's Global Health Institute were among the leading contributors.
Leprosy is one of the oldest recorded and most stigmatized diseases in human history. The disease was prevalent in Europe until the 16th century and is still endemic in many countries, with over 200,000 new cases reported annually. It is caused mainly by the bacterium Mycobacterium leprae, which clusters into several strain types. Only two of these were known to be present in Medieval Europe.
But a new study now shows that there was much more diversity in the leprosy strains circulating in Medieval Europe than previously thought. This finding, based on the sequencing of 10 new ancient genomes from the leprosy-causing bacterium Mycobacterium leprae, complicates prior assumptions about the origin and spread of the disease, and also includes the oldest M. leprae genome sequenced to date, from about 400 AD in the United Kingdom.
Published in PLOS Pathogens, the research was carried out by scientists from EPFL’s Global Health Institute, the Max Planck Institute, the University of Tübingen, the University of Zurich, and others from the United Kingdom, Denmark, Germany, India, the Czech Republic, Hungary, Italy, and France, making it the largest study of its kind.
The scientists wanted to investigate the history and origin of M. leprae further than previous studies. To do this, they looked for genetic evidence from a large number of ancient samples from leprosy throughout Europe.
“Leprosy disappeared from Europe, but it used to be rampant in the middle Ages, and we know very little about why this was case,” says Andrej Benjak from the Global Health Institute at EPFL and one of the co-authors of the study. “But unlike Europe, leprosy is still a problem in many endemic countries. Studying the past spread of M. leprae might help us identify mechanisms that still contribute to the persistence of this disease around the world.”
The scientists examined approximately 90 individuals with skeletal deformations that were characteristic of leprosy. The individuals originated across Europe and from time periods ranging from approximately 400 AD to 1400 AD.
From these samples, the researchers were able to fully reconstruct ten new medieval M. leprae genomes, which represent almost all currently known M. leprae strain types found today at different regions, including Asia, Africa, and the Americas. Additionally, multiple strain types were often found in the same cemetery, illustrating the diversity of the leprosy strains circulating throughout the continent at the time.
“We found much more genetic diversity in ancient Europe than expected,” says Johannes Krause, the study’s senior author and a director at the Max Planck Institute for the Science of Human History. “Additionally, we found that all known strains of leprosy are present in medieval Europe, suggesting that leprosy may already have been widespread throughout Asia and Europe in antiquity or that it might have originated in western Eurasia.”
The oldest leprosy genome
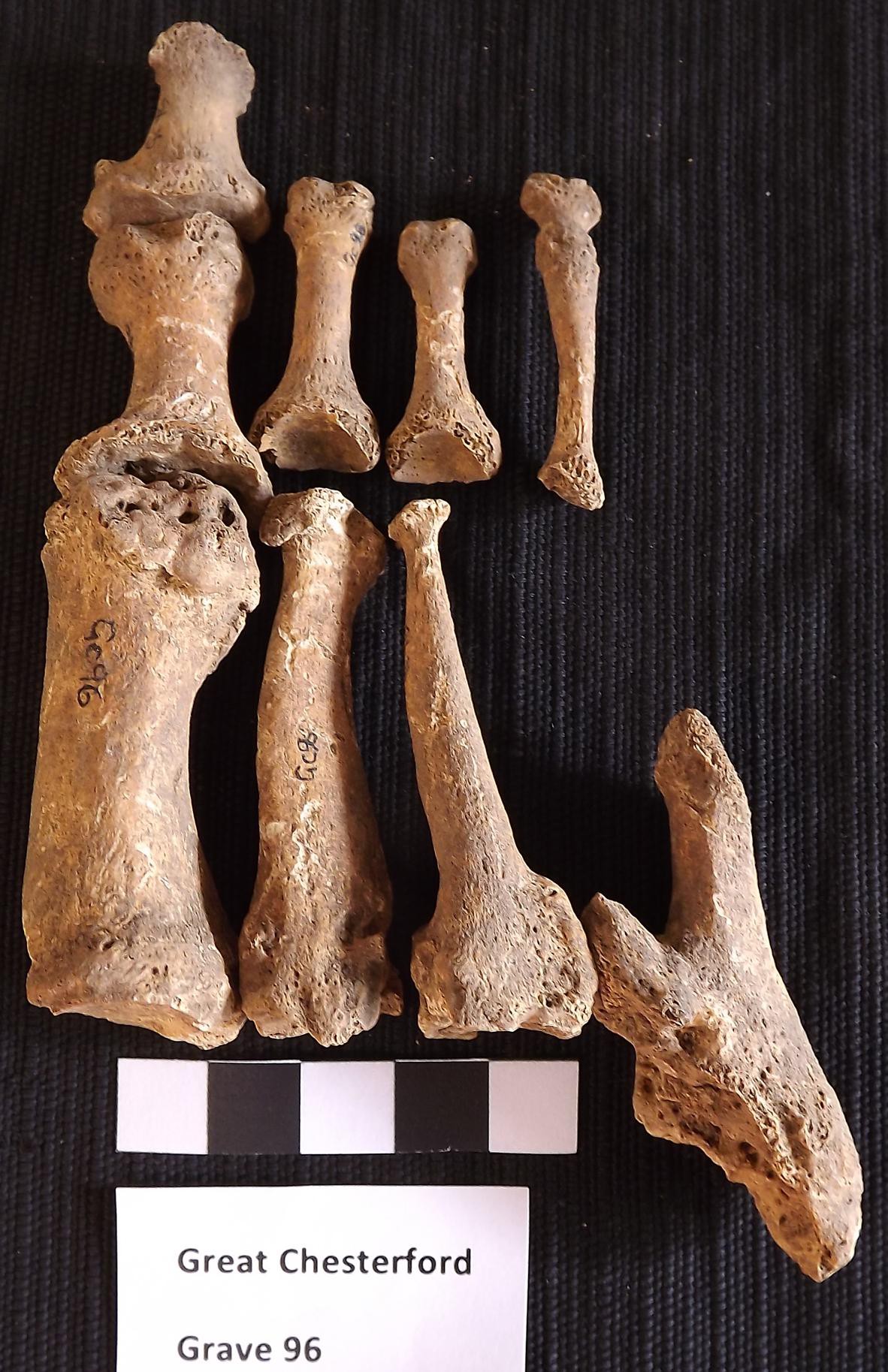
One M. leprae genome reconstructed by the team was from Great Chesterford (United Kingdom) and dates between 415-545 AD. This is the oldest M. leprae genome sequenced to-date, and comes from one of the oldest known cases of leprosy in the United Kingdom. Interestingly, this strain is the same found in modern-day red squirrels and supports the hypothesis that squirrels and the squirrel fur trade were a factor in the spread of leprosy among humans in Europe during the middle ages.
“The dynamics of M. leprae transmission throughout human history are not fully resolved,” says lead author Verena Schuenemann of the University of Zurich. “Characterization and geographic association of the most ancestral strains are crucial for deciphering leprosy’s exact origin. While we have some written records of leprosy cases that predate the Common Era, none of these have yet been confirmed on a molecular level.”
The abundance of ancient genomes in the current study has resulted in an older estimate for the age of M. leprae than previous studies, making it at least a few thousand years old. “Having more ancient genomes in a dating analysis will result in more accurate estimates,” explains Krause. “The next step is to search for even older osteological cases of leprosy than currently available, using well-established methods for identification of potential cases.”
Fondation Raoul Follereau (S.TC.)
Swiss National Science Foundation (SNSF)
European Molecular Biology Organization (EMBO)
DFG Graduate School Human Development in Landscapes Kiel University
European Research Council (ERC)
Max Planck Society
Hungarian National Research, Development and Innovation Office
Verena J. Schuenemann, Charlotte Avanzi, Ben Krause-Kyora, Alexander Seitz, Alexander Herbig, Sarah Inskip, Marion Bonazzi, Ella Reiter, Christian Urban, Dorthe Dangvard Pedersen, G. Michael Taylor, Pushpendra Singh, Graham R. Stewart, Petr Velemínský, Jakub Likovsky, Antónia Marcsik, Erika Molnár, György Pálfi, Valentina Mariotti, Alessandro Riga, M. Giovanna Belcastro, Jesper L. Boldsen, Almut Nebel, Simon Mays, Helen D. Donoghue, Sonia Zakrzewski, Andrej Benjak, Kay Nieselt, Stewart T. Cole, Johannes Krause. Ancient genomes reveal a high diversity of Mycobacterium leprae in medieval Europe. PLOS Pathogens, 14(5): e1006997. DOI: 10.1371/journal.ppat.1006997.