A lung pathogen's dilemma: infect or resist antibiotics?
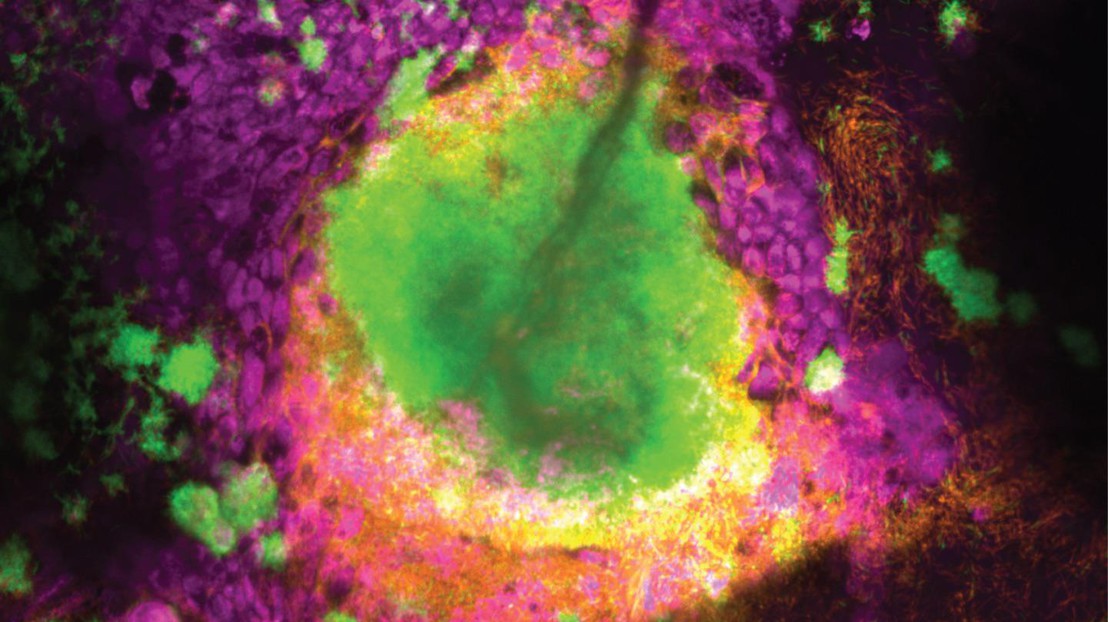
A biofilm formed by two Pseudomonas aeruginosa strains (chronic-like in green, acute-like in orange) growing inside the airway epithelium (magenta). Credit: Lucas Meirelles (EPFL)
A new study by EPFL reveals that the notorious bacterial pathogen Pseudomonas aeruginosa must balance between effectively colonizing human airways and developing antibiotic tolerance to survive.
Imagine trying to settle into a new home while constantly being attacked. That’s what the bacterium Pseudomonas aeruginosa faces when it infects the lungs, and it can’t both spread and protect itself from antibiotics at the same time. Nonetheless, it’s one of the top culprits in hospital-acquired infections and it’s notorious for causing long-lasting, antibiotic-resistant infections, causing damage especially in people with lung diseases like cystic fibrosis, COPD, or bronchiectasis.
To survive tough conditions, P. aeruginosa forms colonies known as “biofilms” – clusters of bacteria encased in a self-produced matrix that provides them with significant advantages, including protection from antibiotics.
But biofilms come at a cost: the clustered bacteria also lose the ability to move around, find nutrients, and spread effectively. For P. aeruginosa infecting a lung, this poses a dilemma: should it spread across the lung’s surface or bunker down to resist incoming antibiotics? Achieving the right balance can mean life and death for the pathogen – and disrupting it can mean life or death for patients.
New research by the group of Alexandre Persat at EPFL’s Global Health Institute has now uncovered how P. aeruginosa manages the trade-off between colonizing and surviving during infection by switching between biofilm formation for antibiotic protection and a more mobile, “planktonic” state to spread and access nutrients, depending on the environmental pressures they face. The study is published in Nature Microbiology.

Mimicking natural infection environments to observe the bacteria
To better understand P. aeruginosa’s behavior, the researchers grew the bacteria on mucus-covered tissue models that mimic human lungs. These tissue models, known as “organoids”, are at the cutting-edge of bioengineering.
“We then used a high throughput screening technique called transposon-insertion sequencing (Tn-seq), combined with metabolic modeling and live imaging, to study how P. aeruginosa adapts to colonize the mucosal surface of the lung and tolerate antibiotic treatment,” says Lucas Meirelles, who led the study.
Thanks to the Tn-seq technique, the scientists identified which genes were important for the bacterium’s survival under different conditions: those which contributed to fitness during mucosal colonization and those which helped the bacteria tolerate antibiotics.
The scientists also used computational modeling to simulate how the bacteria metabolize nutrients in the lung environment, which helped pinpoint the exact metabolic pathways P. aeruginosa relies on during infection.
Finding the right balance
The study found that P. aeruginosa adapts to the lung’s mucus by relying on sugars and lactate, nutrients that are abundant in infected lungs. However, to survive on the mucus, the bacterium also needs to synthesize essential but less available nutrients, like amino acids. This self-sufficiency, or “metabolic independence”, helps the bacterium thrive in the early stages of lung infection.
What Persat's team uncovered is the mechanism behind this dilemma. They found that biofilm formation imposes a “metabolic burden,” meaning that producing the sticky matrix that holds the biofilm together consumes resources, slowing down the bacteria’s ability to spread. In experiments, bacteria that couldn’t form biofilms spread more efficiently but were left vulnerable to antibiotics. This new insight into the metabolic costs of biofilm formation explains how the bacterium balances growth and antibiotic tolerance.
The study highlights the delicate balancing act that Pseudomonas aeruginosa must perform during infections. While the bacteria need to colonize the lung effectively, their best survival strategy – forming biofilms – limits their access to nutrients and, therefore, their ability to spread. However, once antibiotics are introduced, biofilm formation becomes advantageous, protecting the bacteria from being wiped out.
Exploring new paths
The discovery opens the door for the exploration of new treatment strategies: if we can find a way to disrupt the bacteria’s ability to form biofilms without giving them more room to spread, it could make them more vulnerable to existing treatments. And therapies that target the bacteria’s metabolic pathways may also prove to be effective at weakening Pseudomonas infections.
More broadly, the scientists believe that studying pathogens like P. aeruginosa in infection models that replicate the physiology of human tissues is crucial for combating antibiotic resistance.
“Antibiotic resistance is set to become one of the most serious healthcare challenges of this century, and P. aeruginosa is a major contributor to this issue,” says Meirelles. “By using tissue-engineering to replicate the airway environment in the lab, we aim to better understand the physiology of this pathogen. Our hope is that this will uncover previously unknown targets to help us combat these infections and address antibiotic resistance.”
Other contributors
EPFL Laboratory of Computational Systems Biotechnology
Swiss National Science Foundation (SNSF)
NCCR AntiResist
European Molecular Biology Organization (EMBO)
NCCR Microbiomes
Lucas A. Meirelles, Evangelia Vayena, Auriane Debache, Eric Schmidt, Tamara Rossy, Tania Distler, Vassily Hatzimanikatis, Alexandre Persat. Pseudomonas aeruginosa faces a fitness trade-off between mucosal colonization and antibiotic tolerance during airway infections. Nature Microbiology 25 October 2024. DOI: 10.1038/s41564-024-01842-3